anemones
spatstat.data::anemones |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 231 points
# marks are numeric, of storage type 'integer'
# window: rectangle = [0, 280] x [0, 180] unitsChapter 10 demonstrates several, though not all, data objects from package datasets (R version 4.5.3 (2026-03-11)) and package spatstat.data (v3.1.9, GPL (>= 2)).
The function calls in Chapter 10 are exclusively those provided in package base and stats (R version 4.5.3 (2026-03-11)), and in the spatstat.* family of packages.
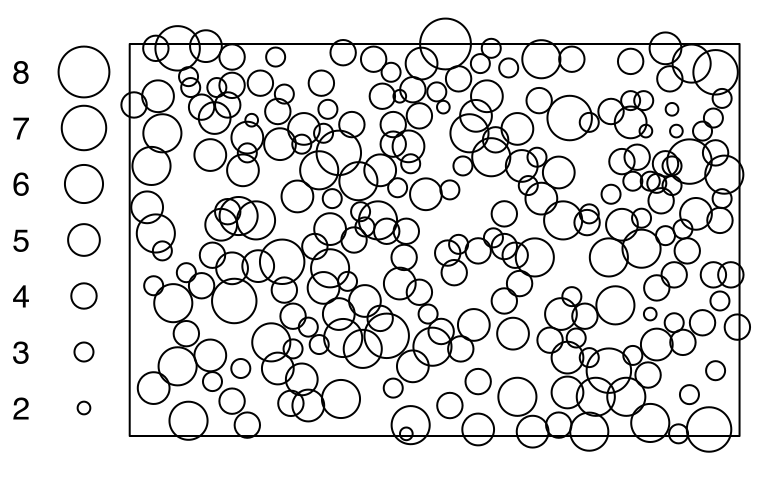
anemonesThe point-pattern (ppp.object, Chapter 36) anemones from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.2, Figure 10.1)
'vector' mark-format (Listing 10.6).anemones
par(mar = c(0,0,0,0))
spatstat.data::anemones |>
spatstat.geom::plot.ppp(main = NULL)anemones
anemones
spatstat.data::anemones |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 231 points
# marks are numeric, of storage type 'integer'
# window: rectangle = [0, 280] x [0, 180] unitsanemones
spatstat.data::anemones |>
spatstat.geom::npoints.ppp()
# [1] 231anemones
spatstat.data::anemones |>
spatstat.geom::Window.ppp()
# window: rectangle = [0, 280] x [0, 180] unitsanemones
spatstat.data::anemones |>
spatstat.geom::marks.ppp() |>
typeof()
# [1] "integer"anemones
spatstat.data::anemones |>
spatstat.geom::markformat.ppp()
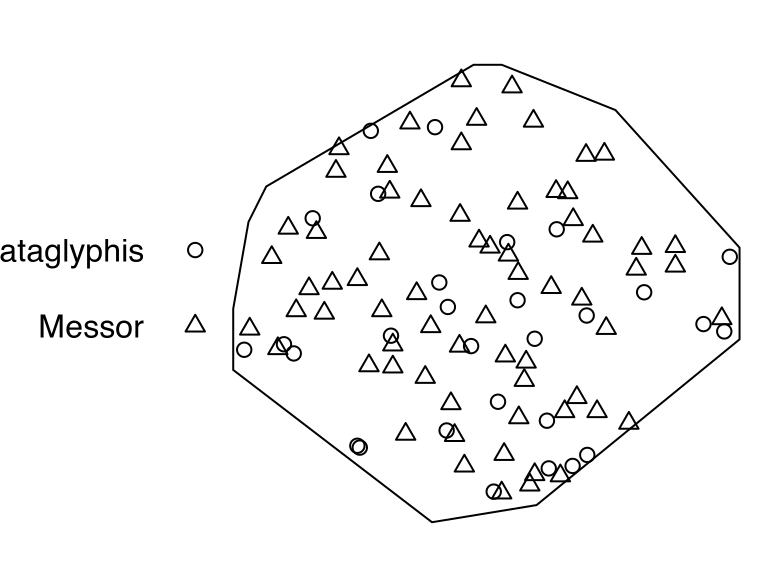
# [1] "vector"antsThe point-pattern (ppp.object, Chapter 36) ants from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.8, Figure 10.2)
'Cataglyphis' and 'Messor' (Listing 10.9);'vector' mark-format.ants
par(mar = c(0,0,0,0))
spatstat.data::ants |>
spatstat.geom::plot.ppp(main = NULL)ants
ants
spatstat.data::ants |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 97 points
# Multitype, with levels = Cataglyphis, Messor
# window: polygonal boundary
# enclosing rectangle: [-25, 803] x [-49, 717] units (one unit = 0.5 feet)ants
spatstat.data::ants |>
spatstat.geom::marks.ppp() |>
table()
#
# Cataglyphis Messor
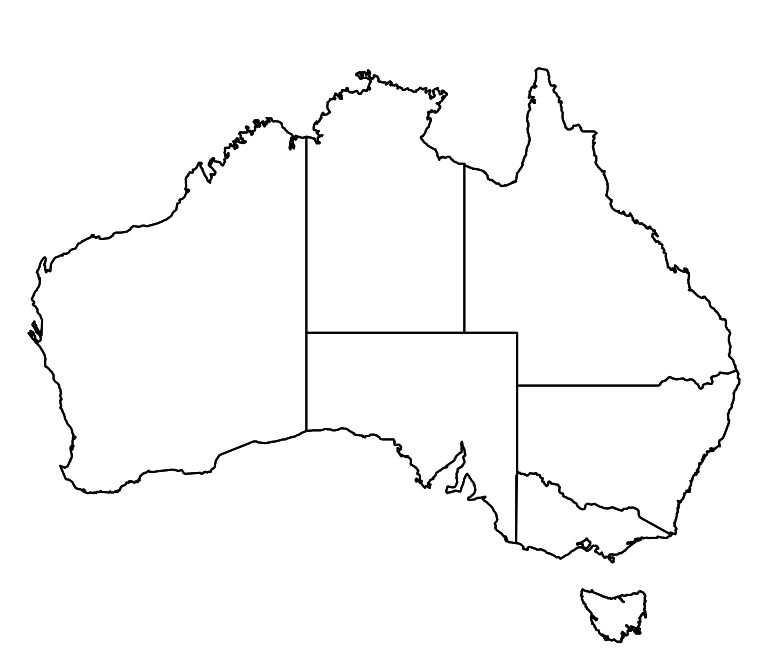
# 29 68austatesThe tessellation (Chapter 42) austates from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.11, Figure 10.3)
austates
par(mar = c(0,0,1,0))
spatstat.data::austates |>
spatstat.geom::plot.tess(main = '')austates
austates
spatstat.data::austates |>
spatstat.geom::print.tess()
# Tessellation
# Tiles are irregular polygons
# 7 tiles (irregular windows)
# window: polygonal boundary
# enclosing rectangle: [113.19392, 153.6692] x [-43.59316, -10.93156] degreesaustates
spatstat.data::austates |>
spatstat.geom::tiles()
# List of spatial objects
#
# WA:
# window: polygonal boundary
# enclosing rectangle: [113.19392, 129.01141] x [-35.11407, -13.76426] degrees
#
# NT:
# window: polygonal boundary
# enclosing rectangle: [129.01141, 138.0038] x [-25.988593, -11.045627] degrees
#
# ✂️ --- output truncated --- ✂️betacellsThe point-pattern (ppp.object, Chapter 36) betacells from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.14, Figure 10.4)
'dataframe' mark-format (Listing 10.15);area (Listing 10.16);type with two levels, 'off' and 'on' (Listing 10.16).betacells
par(mar = c(0,0,0,0))
spatstat.data::betacells |>
spatstat.geom::plot.ppp(main = '')betacells
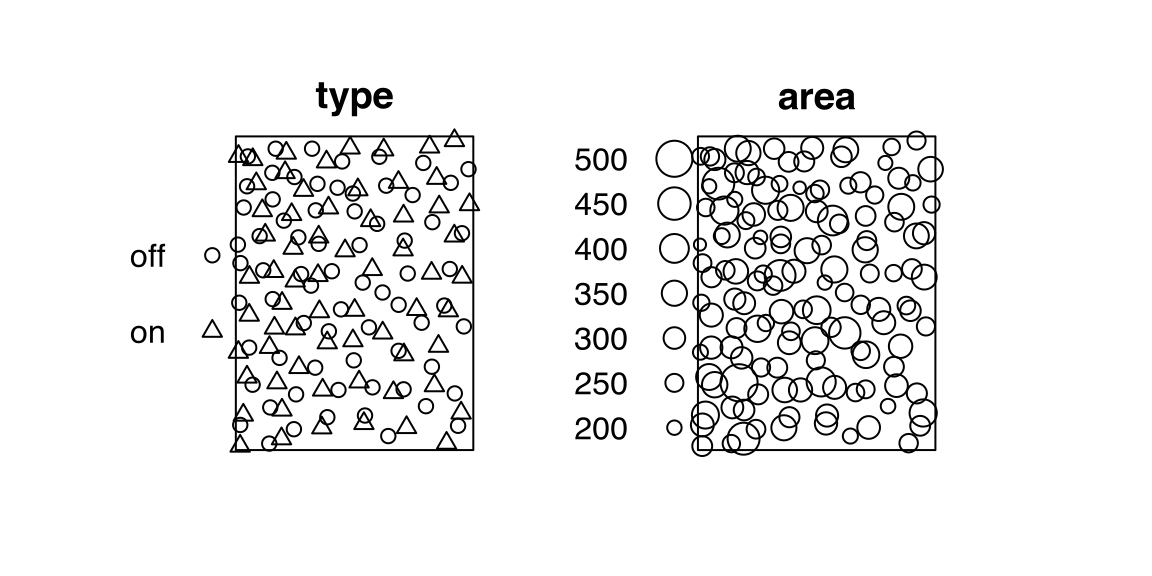

betacells
spatstat.data::betacells |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 135 points
# Mark variables: type, area
# window: rectangle = [28.08, 778.08] x [16.2, 1007.02] micronsbetacells
spatstat.data::betacells |>
spatstat.geom::markformat.ppp()
# [1] "dataframe"betacells
spatstat.data::betacells |>
spatstat.geom::marks.ppp()
# type area
# 1 on 275.9
# 2 off 241.2
# 3 on 256.0
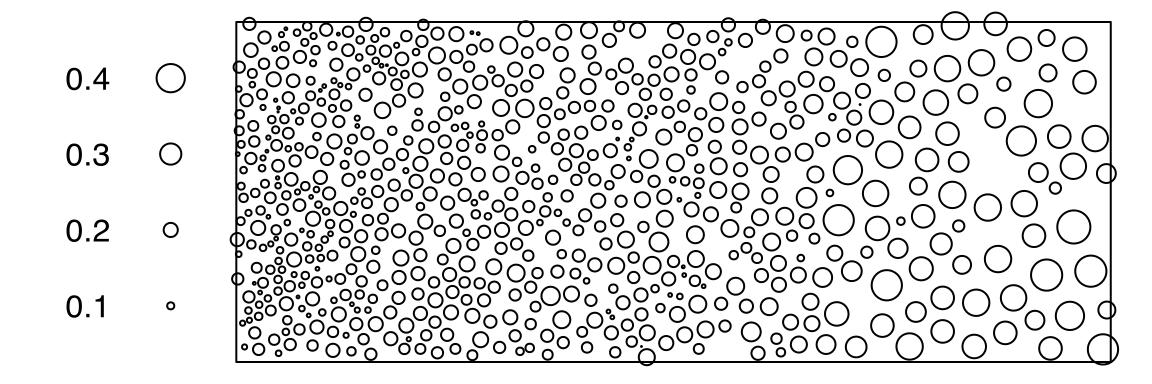
# ✂️ --- output truncated --- ✂️bronzefilterThe point-pattern (ppp.object, Chapter 36) bronzefilter from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.18, Figure 10.5)
bronzefilter
par(mar = c(0,1,0,0))
spatstat.data::bronzefilter |>
spatstat.geom::plot.ppp(main = NULL)bronzefilter
bronzefilter
spatstat.data::bronzefilter |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 678 points
# marks are numeric, of storage type 'double'
# window: rectangle = [0, 18] x [0, 7] mmbtb.extraThe point-pattern-list (ppplist, Chapter 37) btb.extra from package spatstat.data (v3.1.9, GPL (>= 2)) (Listing 10.20, Figure 10.6)
S3 class 'solist' (Chapter 39, Listing 10.21);ppp.object, Chapter 36) members (Listing 10.22).btb.extra
par(mar = c(0,1,1,1))
spatstat.data::btb.extra |>
spatstat.geom::plot.solist()btb.extra
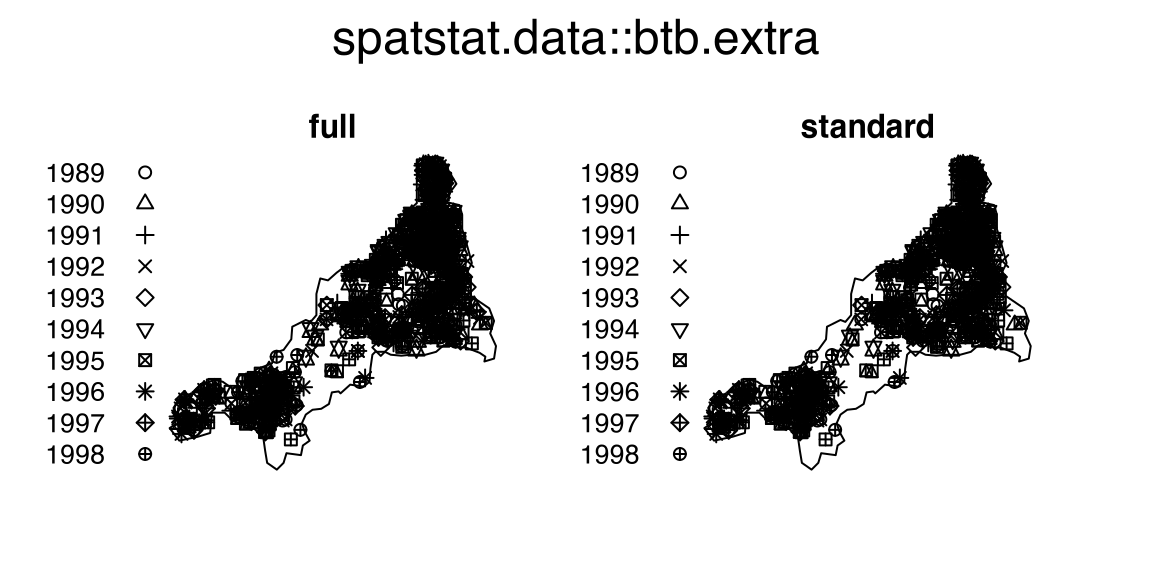
btb.extra
spatstat.data::btb.extra
# List of point patterns
#
# full:
# Marked planar point pattern: 919 points
# Mark variables: year, spoligotype
# window: polygonal boundary
# enclosing rectangle: [133.5147, 246.0193] x [10.88514, 118.7298] km
#
# standard:
# Marked planar point pattern: 873 points
# Mark variables: year, spoligotype
# window: polygonal boundary
# enclosing rectangle: [133.5147, 246.0193] x [10.88514, 118.7298] kmbtb.extra
spatstat.data::btb.extra |>
class()
# [1] "ppplist" "solist" "anylist" "listof" "list"class of members of btb.extra
spatstat.data::btb.extra |>
sapply(FUN = class)
# full standard
# "ppp" "ppp"carsThe data frame (data.frame, Chapter 18) cars from package datasets (R version 4.5.3 (2026-03-11)) has (Listing 10.23)
$speed and $dist.cars
datasets::cars |>
print.data.frame()
# speed dist
# 1 4 2
# 2 4 10
# 3 7 4
# 4 7 22
# ✂️ --- output truncated --- ✂️dimensions of cars
datasets::cars |>
dim.data.frame()
# [1] 50 2cetaceansThe hyper data frame (hyperframe, Chapter 26) cetaceans from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.25)
ppp, Chapter 36) hypercolumns: $whales, $dolphins, $fish and $plankton.cetaceans
spatstat.data::cetaceans |>
spatstat.geom::print.hyperframe()
# Hyperframe:
# whales dolphins fish plankton
# 1 (ppp) (ppp) (ppp) (ppp)
# 2 (ppp) (ppp) (ppp) (ppp)
# 3 (ppp) (ppp) (ppp) (ppp)
# ✂️ --- output truncated --- ✂️dimensions of cetaceans
spatstat.data::cetaceans |>
spatstat.geom::dim.hyperframe()
# [1] 9 4demohyperThe hyper data frame (hyperframe, Chapter 26) demohyper from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.27)
ppp, Chapter 36) hypercolumn $Pointsim, Chapter 28) hypercolumn $Image$Group.demohyper
spatstat.data::demohyper |>
spatstat.geom::print.hyperframe()
# Hyperframe:
# Points Image Group
# 1 (ppp) (im) a
# 2 (ppp) (im) b
# 3 (ppp) (im) adimensions of demohyper
spatstat.data::demohyper |>
spatstat.geom::dim.hyperframe()
# [1] 3 3To view the hyper data frame demohyper in a desired format, readers may call the S3 method spatstat.geom::print.hyperframe() explicitly (Listing 10.27). Alternatively, readers may call the S3 generic function print() by simply typing demohyper at the R console prompt and pressing Enter, after putting the package spatstat.geom (v3.7.3, GPL (>= 2))
search() path, by either one of the following approaches,
library(), e.g., library(spatstat.geom), which is called internally by the function require();attachNamespace(), e.g., attachNamespace('spatstat.geom');loadedNamespaces(), by either one of the following approaches,
loadNamespace(), e.g., loadNamespace('spatstat.geom'), which is called internally by the function requireNamespace();spatstat.geom (v3.7.3, GPL (>= 2)) explicitly with its namespace, e.g., spatstat.geom::dim.hyperframe to print the function itself, or Listing 10.28, Listing 10.29, etc.The rest of Section 10.9 showcases the *.hyperframe() methods of the .Primitive S3 generic functions names() (Listing 10.29) and `$` (Listing 10.30, Listing 10.31).
Listing 10.29 finds the (hyper)column names of the hyper data frame demohyper,
demohyper
spatstat.data::demohyper |>
spatstat.geom::names.hyperframe()
# [1] "Points" "Image" "Group"Listing 10.30 and Listing 10.31 observe the ppp-hypercolumn $Points,
ppp-hypercolumn $Points
spatstat.data::demohyper$Points |>
class()
# [1] "ppplist" "solist" "anylist" "listof" "list"ppp-hypercolumn $Points, nerdy!
spatstat.data::demohyper |>
spatstat.geom::`$.hyperframe`(name = 'Points') |> # nerdy!!
identical(y = spatstat.data::demohyper$Points) |>
stopifnot()Listing 10.32 and Listing 10.33 find the first point-pattern element of the ppp-hypercolumn $Points,
ppp-hypercolumn $Points
demohyper_p1 = spatstat.data::demohyper$Points[[1L]]
demohyper_p1 |>
spatstat.geom::print.ppp()
# Planar point pattern: 104 points
# window: binary image mask
# 128 x 128 pixel array (ny, nx)
# enclosing rectangle: [2.017, 3.93] x [0.645, 3.278] unitsppp-hypercolumn $Points, nerdy!
spatstat.data::demohyper$Points |>
base::`[[`(i = 1L) |> # nerdy!!
identical(y = demohyper_p1) |>
stopifnot()Listing 10.34 finds the first pixel-image element of the im-hypercolumn $Image,
im-hypercolumn $Image
spatstat.data::demohyper$Image[[1L]] |>
spatstat.geom::print.im()
# real-valued pixel image
# 53 x 39 pixel array (ny, nx)
# enclosing rectangle: [2.017, 3.93] x [0.645, 3.278] unitsfaithfulThe data frame (data.frame, Chapter 18) faithful from package datasets (R version 4.5.3 (2026-03-11)) has (Listing 10.35)
$eruptions and $waiting.faithful
datasets::faithful |>
print.data.frame()
# eruptions waiting
# 1 3.600 79
# 2 1.800 54
# 3 3.333 74
# 4 2.283 62
# ✂️ --- output truncated --- ✂️dimensions of faithful
datasets::faithful |>
dim.data.frame()
# [1] 272 2finpinesThe point-pattern (ppp.object, Chapter 36) finpines from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.38, Figure 10.7)
diameter and height.finpines
par(mar = c(0,0,0,0))
spatstat.data::finpines |>
spatstat.geom::plot.ppp(main = '')finpines
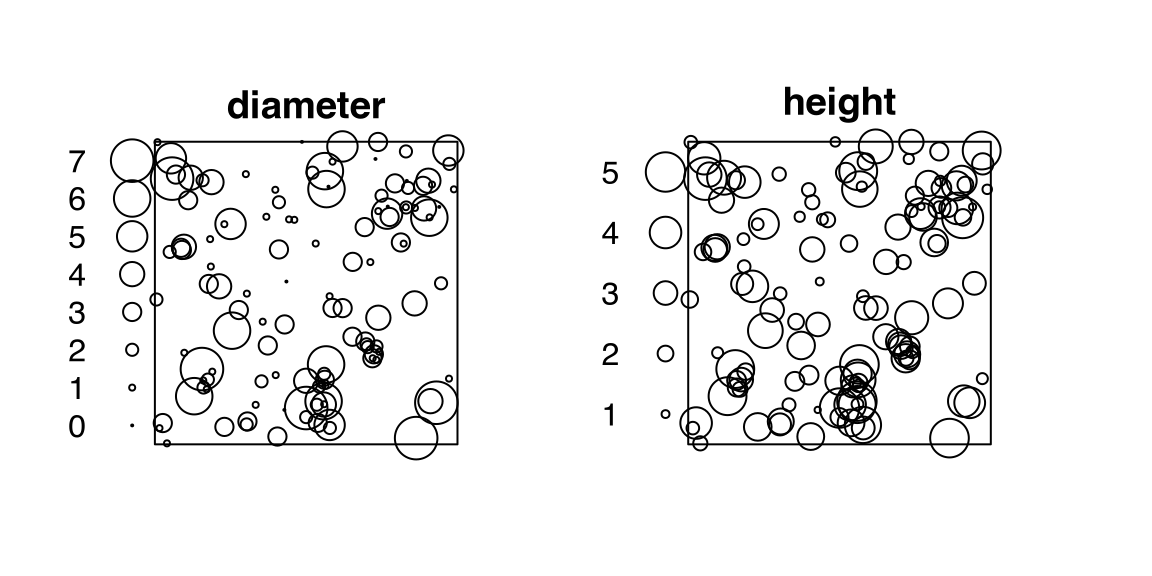
finpines
spatstat.data::finpines |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 126 points
# Mark variables: diameter, height
# window: rectangle = [-5, 5] x [-8, 2] metresfluThe hyper data frame (hyperframe, Chapter 26) flu from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.39)
ppp, Chapter 36) hypercolumn $pattern$virustype, $stain, $frameidflu
spatstat.data::flu |>
spatstat.geom::print.hyperframe()
# Hyperframe:
# pattern virustype stain frameid
# wt M2-M1 13 (ppp) wt M2-M1 13
# wt M2-M1 22 (ppp) wt M2-M1 22
# wt M2-M1 27 (ppp) wt M2-M1 27
# ✂️ --- output truncated --- ✂️dimensions of flu
spatstat.data::flu |>
spatstat.geom::dim.hyperframe()
# [1] 41 4gorillasThe point-pattern (ppp.object, Chapter 36) gorillas from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.42, Figure 10.8)
group (with two levels 'major' and 'minor') and season (with two levels 'dry' and 'rainy').gorillas
par(mar = c(0,0,1,0))
spatstat.data::gorillas |>
spatstat.geom::plot.ppp(which.marks = c('group', 'season'))gorillas
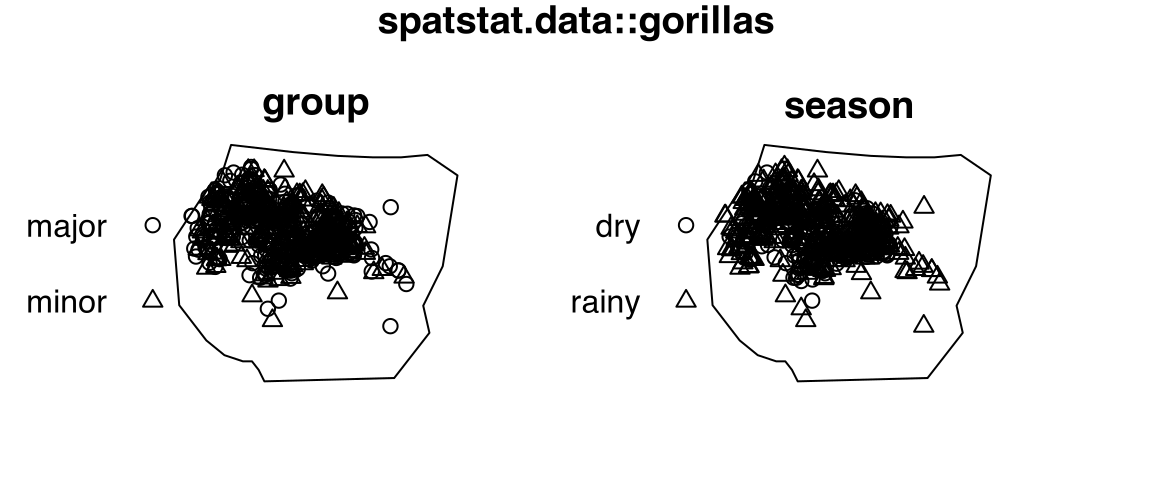
gorillas
spatstat.data::gorillas |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 647 points
# Mark variables: group, season, date
# window: polygonal boundary
# enclosing rectangle: [580457.9, 585934] x [674172.8, 678739.2] metresgorillas.extraThe pixel-image list (imlist, Chapter 29) gorillas.extra from package spatstat.data (v3.1.9, GPL (>= 2)) (Listing 10.44, Figure 10.9)
S3 class 'solist' (Chapter 39, Listing 10.45);im, Chapter 28) members (Listing 10.46).gorillas.extra
par(mar = c(0,0,0,0))
spatstat.data::gorillas.extra |>
plot(main = '')gorillas.extra
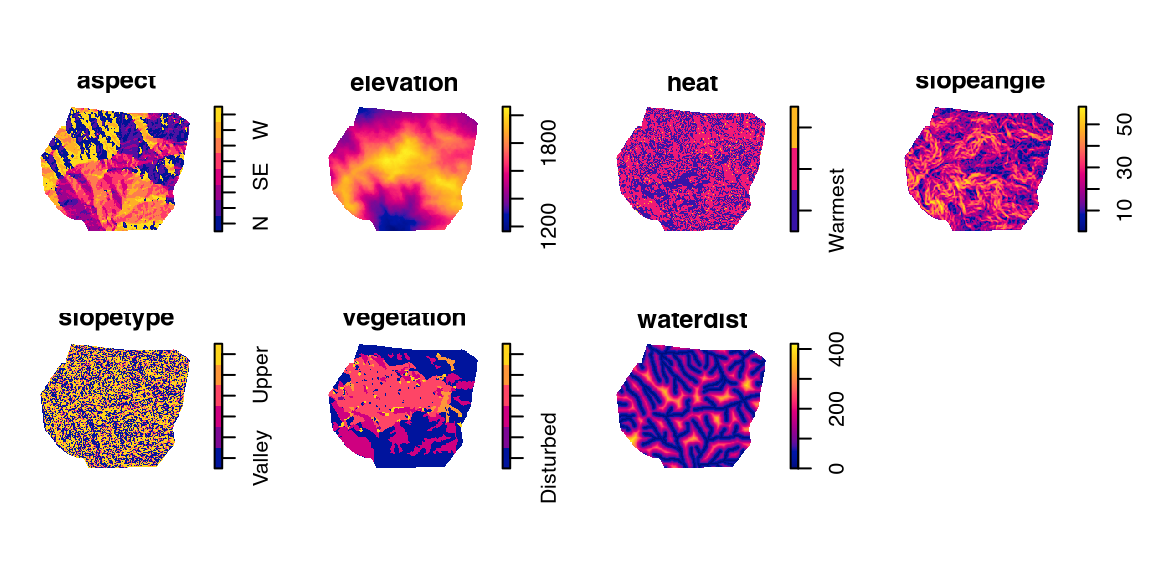
gorillas.extra
spatstat.data::gorillas.extra
# List of pixel images
#
# aspect:
# factor-valued pixel image
# factor levels:
# [1] "N" "NE" "E" "SE" "S" "SW" "W" "NW"
# 149 x 181 pixel array (ny, nx)
# enclosing rectangle: [580440, 586000] x [674160, 678730] metres
#
# elevation:
# integer-valued pixel image
# 149 x 181 pixel array (ny, nx)
# enclosing rectangle: [580440, 586000] x [674160, 678730] metres
#
# ✂️ --- output truncated --- ✂️gorillas.extra
spatstat.data::gorillas.extra |>
class()
# [1] "imlist" "solist" "anylist" "listof" "list"class of members of gorillas.extra
spatstat.data::gorillas.extra |>
sapply(FUN = class)
# aspect elevation heat slopeangle slopetype vegetation waterdist
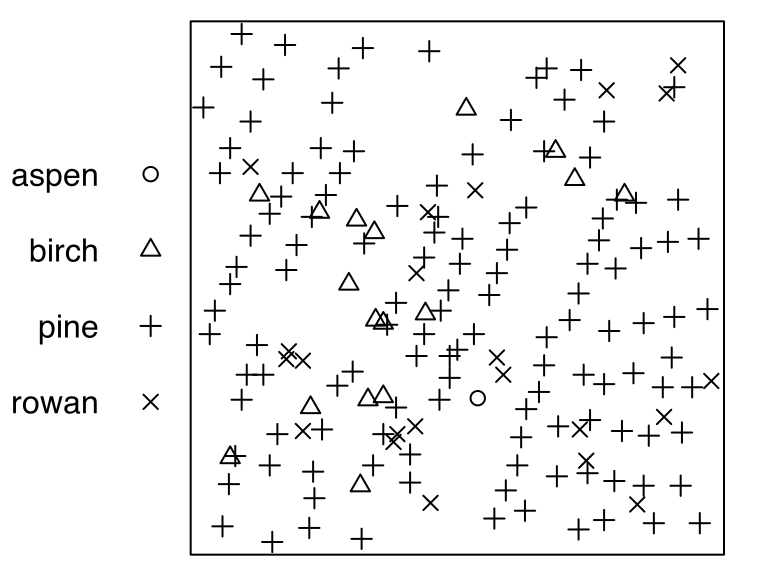
# "im" "im" "im" "im" "im" "im" "im"hyytialaThe point-pattern (ppp.object, Chapter 36) hyytiala from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.48, Figure 10.10)
'aspen', 'birch', 'pine' and 'rowan' (Listing 10.49);'vector' mark-format.hyytiala
par(mar = c(0,0,0,0))
spatstat.data::hyytiala |>
spatstat.geom::plot.ppp(main = NULL)hyytiala
hyytiala
spatstat.data::hyytiala |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 168 points
# Multitype, with levels = aspen, birch, pine, rowan
# window: rectangle = [0, 20] x [0, 20] metreshyytiala
spatstat.data::hyytiala |>
spatstat.geom::marks.ppp() |>
table()
#
# aspen birch pine rowan
# 1 17 128 22KovesiThe hyper data frame (hyperframe, Chapter 26) Kovesi from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.50)
$linear, $diverging, etc.character hypercolumn $values. This is a length-41 (Listing 10.53) anylist (Chapter 15, Listing 10.52) of character vectors, each of them has a length of 256 (Listing 10.54).Kovesi
spatstat.data::Kovesi |>
spatstat.geom::print.hyperframe()
# Hyperframe:
# linear diverging rainbow cyclic isoluminant ternary colsig l1 l2 chro n cycsh values
# 1 FALSE FALSE FALSE TRUE FALSE FALSE j 15 85 0 256 0 (character)
# 2 FALSE FALSE FALSE TRUE FALSE FALSE j 15 85 0 256 25 (character)
# 3 FALSE FALSE FALSE TRUE FALSE FALSE mrybm 35 75 68 256 0 (character)
# 4 FALSE FALSE FALSE TRUE FALSE FALSE mrybm 35 75 68 256 25 (character)
# 5 FALSE FALSE FALSE TRUE FALSE FALSE mygbm 30 95 78 256 0 (character)
# ✂️ --- output truncated --- ✂️dimensions of Kovesi
spatstat.data::Kovesi |>
spatstat.geom::dim.hyperframe()
# [1] 41 13$values
spatstat.data::Kovesi$values |>
class()
# [1] "anylist" "listof" "list"$values
spatstat.data::Kovesi$values |>
length()
# [1] 41$values
spatstat.data::Kovesi$values |>
lengths() |>
unique.default()
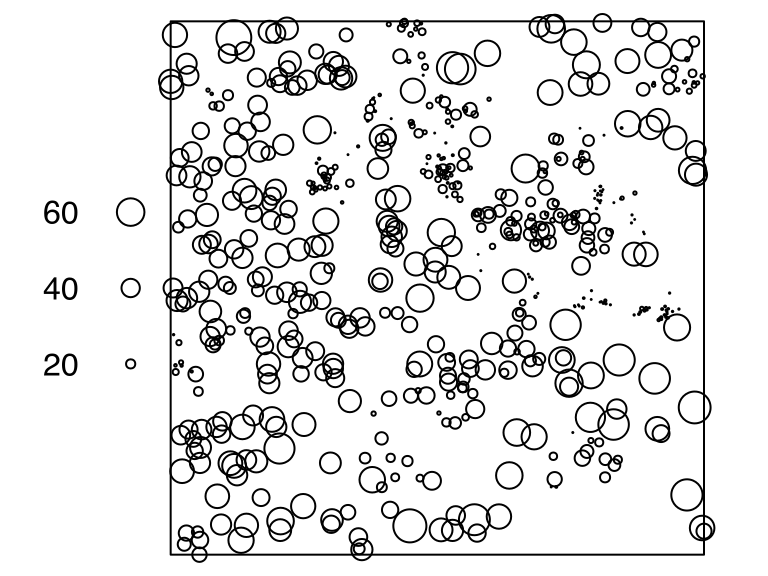
# [1] 256longleafThe point-pattern (ppp.object, Chapter 36) longleaf from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.56, Figure 10.11)
longleaf
par(mar = c(0,0,0,0))
spatstat.data::longleaf |>
spatstat.geom::plot.ppp(main = NULL)longleaf
longleaf
spatstat.data::longleaf |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 584 points
# marks are numeric, of storage type 'double'
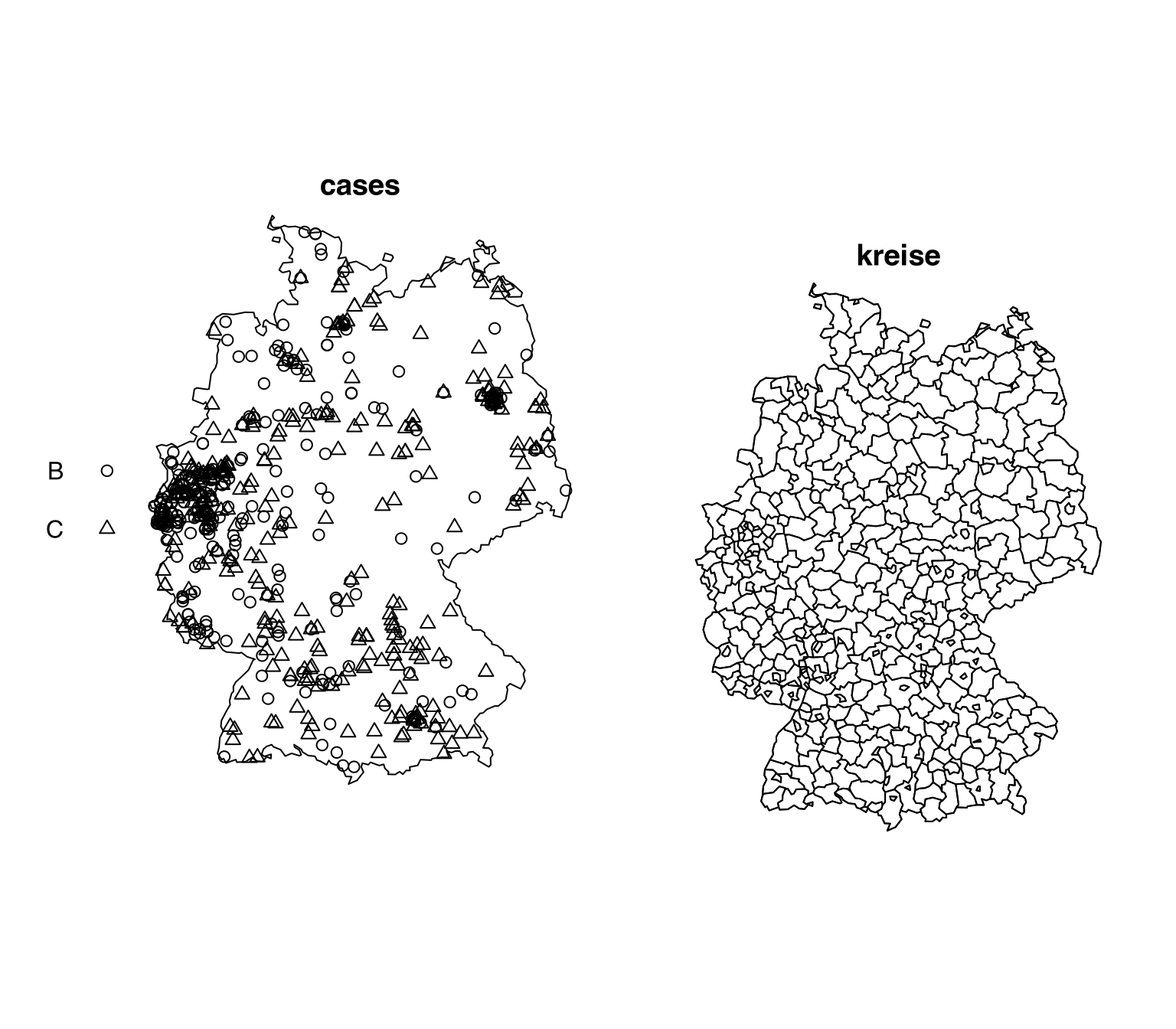
# window: rectangle = [0, 200] x [0, 200] metresmeningitisThe spatial-object list (solist, Chapter 39) meningitis from package spatstat.data (v3.1.9, GPL (>= 2)) contains (Listing 10.58, Figure 10.12)
ppp.object, Chapter 36) $cases;tessellation (Chapter 42) $kreise.meningitis
par(mar = c(0,0,0,0))
spatstat.data::meningitis |>
spatstat.geom::plot.solist(main = '')meningitis
meningitis
spatstat.data::meningitis
# List of spatial objects
#
# cases:
# Marked planar point pattern: 636 points
# Multitype, with levels = B, C
# window: polygonal boundary
# enclosing rectangle: [4031.295, 4672.253] x [2684.102, 3549.931] km
#
# kreise:
# Tessellation
# Tiles are irregular polygons
# 413 tiles (irregular windows)
# Tessellation has a data frame of marks:
# $marks: double
# window: polygonal boundary
# enclosing rectangle: [4031.295, 4672.253] x [2684.102, 3549.931] kmnbfiresThe point-pattern (ppp.object, Chapter 36) nbfires from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.60, Figure 10.13)
$fire.type, $cause and $ign.src;$fnl.size.nbfires
par(mar = c(0,0,1,0))
spatstat.data::nbfires |>
spatstat.geom::plot.ppp(which.marks = c('fire.type', 'cause', 'ign.src', 'fnl.size'))
# Warning: Only 10 out of 16 symbols are shown in the symbol mapnbfires
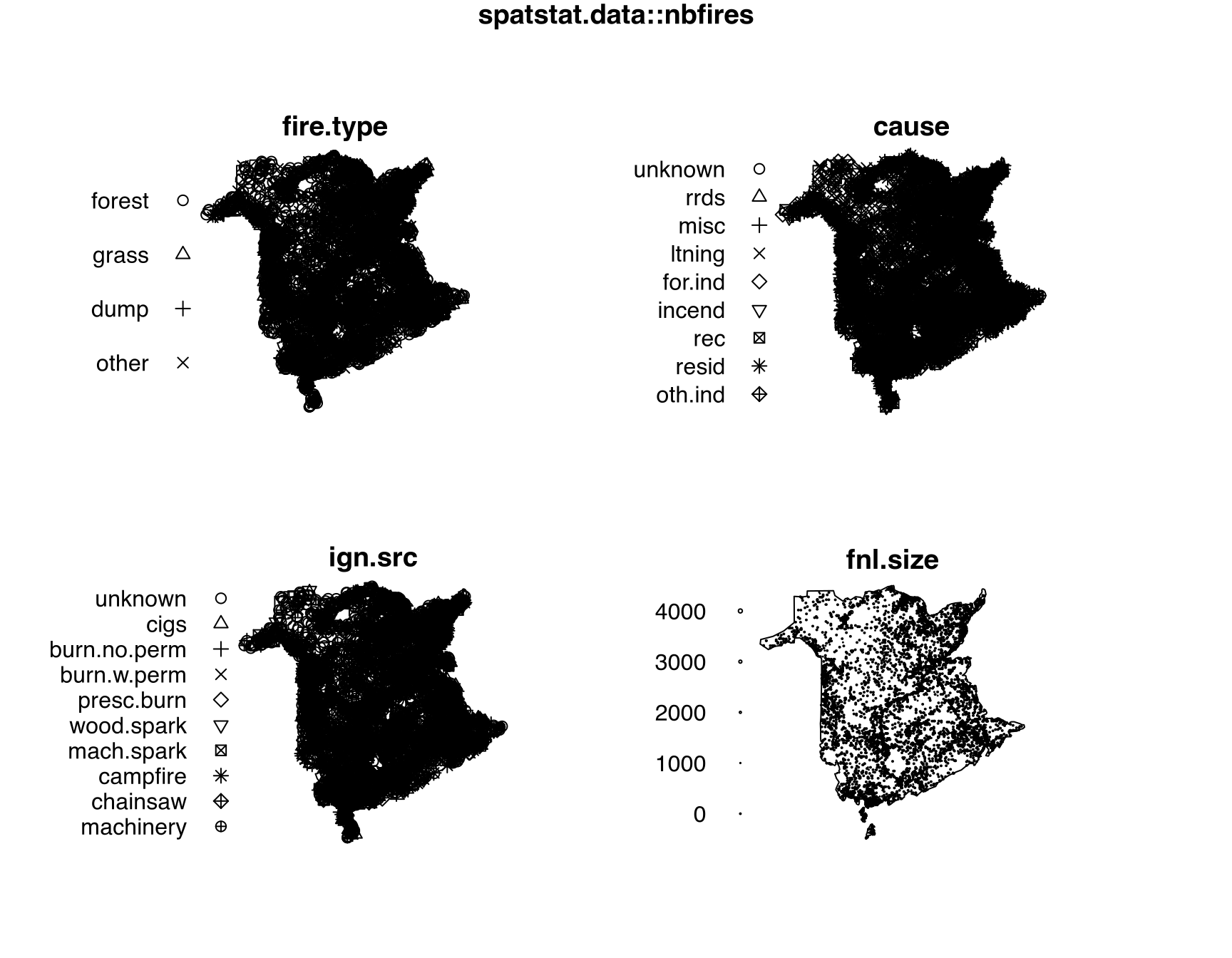

nbfires
spatstat.data::nbfires |>
spatstat.geom::print.ppp()
# Warning: some mark values are NA in the point pattern x
# Marked planar point pattern: 7108 points
# Mark variables: year, fire.type, dis.date, dis.julian, out.date, out.julian, cause, ign.src, fnl.size
# window: polygonal boundary
# enclosing rectangle: [0, 1000] x [0, 958.9142] units (one unit = 0.403716 km)osteoThe hyper data frame (hyperframe, Chapter 26) osteo from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.61)
$brick nested in the bone sample $idpp3, Chapter 35) hypercolumn $ptsosteo
spatstat.data::osteo |>
spatstat.geom::print.hyperframe()
# Hyperframe:
# id shortid brick pts depth
# 1 c77za4 4 1 (pp3) 45
# 2 c77za4 4 2 (pp3) 60
# 3 c77za4 4 3 (pp3) 55
# 4 c77za4 4 4 (pp3) 60
# 5 c77za4 4 5 (pp3) 85
# ✂️ --- output truncated --- ✂️dimensions of osteo
spatstat.data::osteo |>
spatstat.geom::dim.hyperframe()
# [1] 40 5sprucesThe point-pattern (ppp.object, Chapter 36) spruces from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.64, Figure 10.14)
spruces
par(mar = c(0,0,0,0))
spatstat.data::spruces |>
spatstat.geom::plot.ppp(main = NULL)spruces
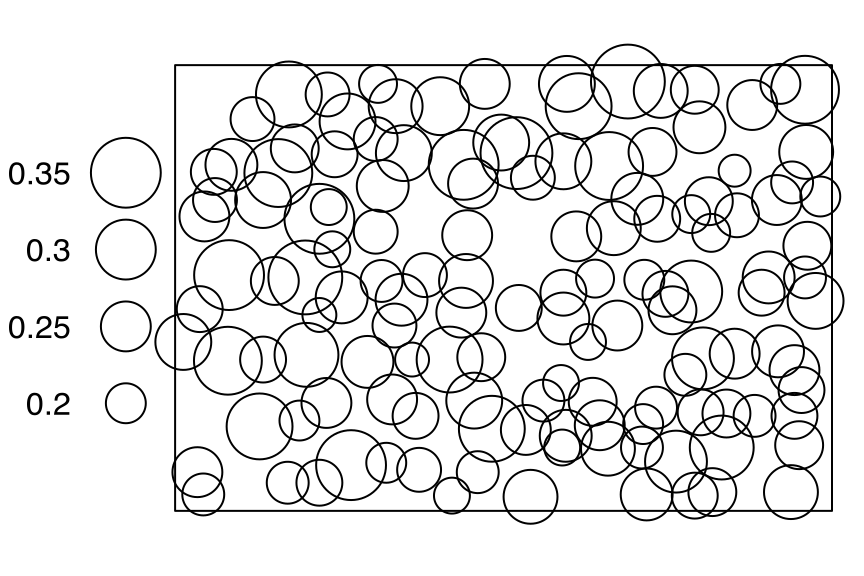

spruces
spatstat.data::spruces |>
spatstat.geom::print.ppp()
# Marked planar point pattern: 134 points
# marks are numeric, of storage type 'double'
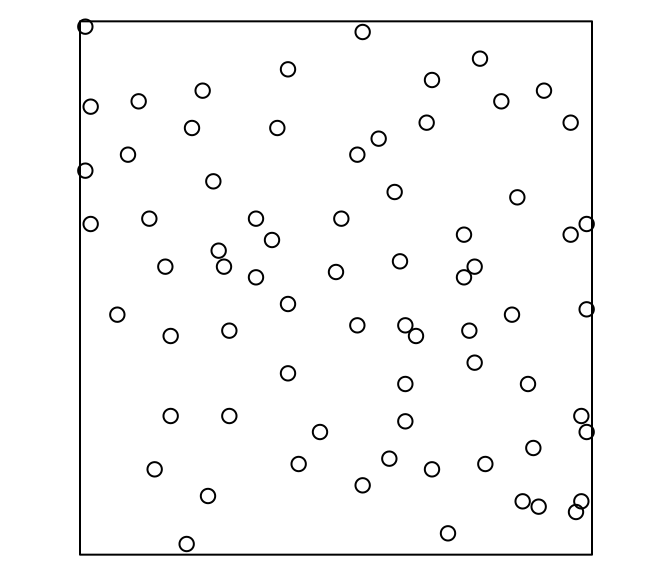
# window: rectangle = [0, 56] x [0, 38] metresswedishpinesThe point-pattern (ppp.object, Chapter 36) swedishpines from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.66, Figure 10.15)
'none' mark-format.swedishpines
par(mar = c(0,0,0,0))
spatstat.data::swedishpines |>
spatstat.geom::plot.ppp(main = NULL)swedishpines
swedishpines
spatstat.data::swedishpines |>
spatstat.geom::print.ppp()
# Planar point pattern: 71 points
# window: rectangle = [0, 96] x [0, 100] units (one unit = 0.1 metres)VADeathsThe matrix VADeaths from package datasets (R version 4.5.3 (2026-03-11)) has (Listing 10.67)
VADeaths
datasets::VADeaths |>
print.default()
# Rural Male Rural Female Urban Male Urban Female
# 50-54 11.7 8.7 15.4 8.4
# 55-59 18.1 11.7 24.3 13.6
# 60-64 26.9 20.3 37.0 19.3
# 65-69 41.0 30.9 54.6 35.1
# 70-74 66.0 54.3 71.1 50.0dimensions of VADeaths
datasets::VADeaths |>
dim()
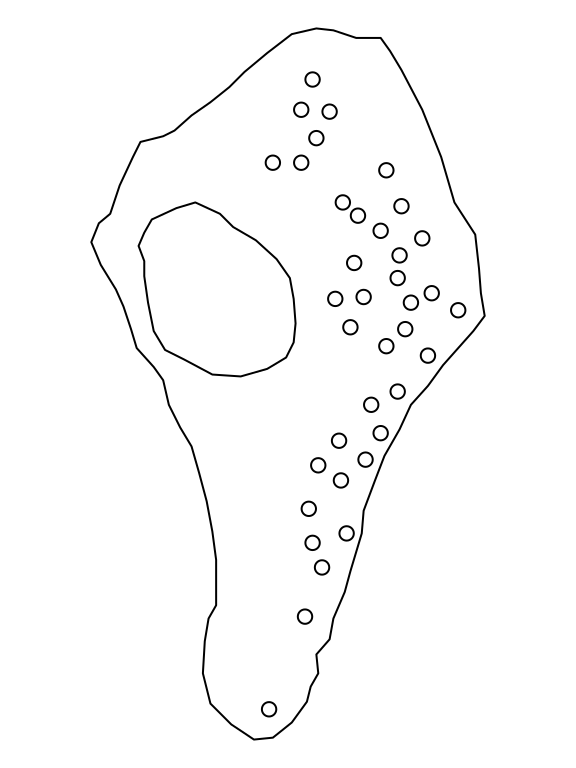
# [1] 5 4vesiclesThe point-pattern (ppp.object, Chapter 36) vesicles from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.70, Figure 10.16)
'none' mark-format.vesicles
par(mar = c(0,0,0,0))
spatstat.data::vesicles |>
spatstat.geom::plot.ppp(main = NULL)vesicles
vesicles
spatstat.data::vesicles |>
spatstat.geom::print.ppp()
# Planar point pattern: 37 points
# window: polygonal boundary
# enclosing rectangle: [22.6796, 586.2292] x [11.9756, 1030.7] nmvesicles.extraThe spatial-object list (solist, Chapter 39) vesicles.extra from package spatstat.data (v3.1.9, GPL (>= 2)) has (Listing 10.71, Listing 10.72)
psp, Chapter 38) $activezone$mitochondria, $presynapse and $maskvesicles.extra
spatstat.data::vesicles.extra
# List of spatial objects
#
# activezone:
# planar line segment pattern: 9 line segments
# window: rectangle = [0, 625] x [0, 1050] nm
#
# mitochondria:
# window: polygonal boundary
# enclosing rectangle: [90.41389, 315.29187] x [532.1753, 781.4376] nm
#
# presynapse:
# window: polygonal boundary
# enclosing rectangle: [22.6796, 586.2292] x [11.9756, 1030.7] nm
#
# mask:
# window: binary image mask
# 420 x 250 pixel array (ny, nx)
# enclosing rectangle: [0, 250] x [0, 420] unitsclass of members of vesicles.extra
spatstat.data::vesicles.extra |>
lapply(FUN = class)
# $activezone
# [1] "psp" "list"
#
# $mitochondria
# [1] "owin"
#
# $presynapse
# [1] "owin"
#
# $mask
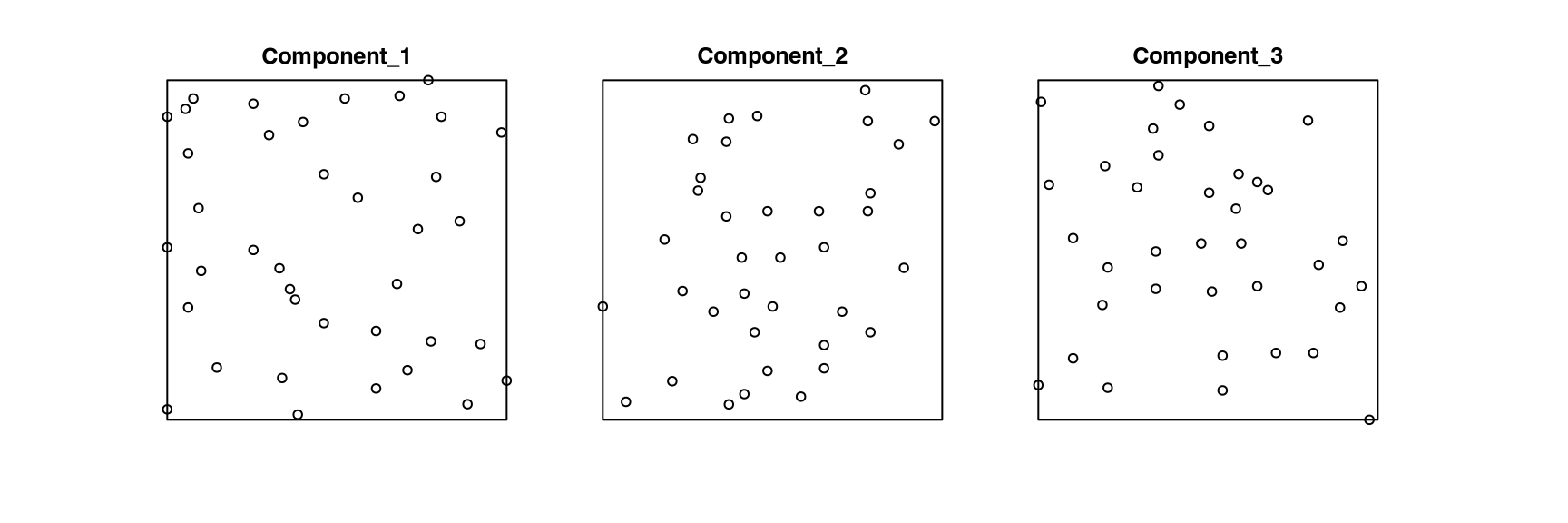
# [1] "owin"waterstridersThe point-pattern-list (ppplist, Chapter 37) waterstriders from package spatstat.data (v3.1.9, GPL (>= 2)) (Listing 10.74, Figure 10.17)
S3 class 'solist' (Chapter 39)ppp.object, Chapter 36) members (Listing 10.75).waterstriders
par(mar = c(0,0,0,0))
spatstat.data::waterstriders |>
spatstat.geom::plot.solist(main = '')waterstriders
waterstriders
spatstat.data::waterstriders
# List of point patterns
#
# Component 1:
# Planar point pattern: 38 points
# window: rectangle = [0, 48.1] x [0, 48.1] cm
#
# Component 2:
# Planar point pattern: 36 points
# window: rectangle = [0, 48.8] x [0, 48.8] cm
#
# Component 3:
# Planar point pattern: 36 points
# window: rectangle = [0, 46.4] x [0, 46.4] cmclass of members of waterstriders
spatstat.data::waterstriders |>
sapply(FUN = class)
# [1] "ppp" "ppp" "ppp"